I have the following code which will generate two pdf files containing plots to the current working directory:
library(reshape)
library(ggplot2)
require(ggplot2)
source("http://gridextra.googlecode.com/svn/trunk/R/arrange.r")
data<-structure(list(Loci = structure(c(1L, 2L, 3L, 4L, 5L, 6L, 7L,
8L, 9L, 10L, 11L, 12L, 13L, 14L, 15L, 16L, 17L, 18L, 19L, 20L,
21L, 22L, 23L, 24L, 25L, 26L, 27L, 28L, 29L, 30L, 31L, 32L, 33L,
34L, 35L, 36L, 37L, 38L, 39L, 40L, 41L, 42L, 43L, 52L, 44L, 45L,
46L, 47L, 48L, 49L, 50L, 51L, 53L, 54L, 55L, 56L, 57L, 58L, 59L,
60L, 61L, 62L), .Label = c("Baez", "Blue", "C147", "C204", "C21",
"C278_PT", "C294", "C316", "C485", "C487_PigTa", "C536", "Carey",
"Cool", "Coyote", "Deadpool", "Epstein", "Glass", "Harrison",
"Harvest", "Hazel", "i-113", "i-114", "i-120_PigT", "i-126",
"i-127", "Imagine", "Jackstraw_", "Jericho", "Jerry-Garc", "Jude",
"Kitty", "Majesty", "Million", "Monkey", "Mozambique", "Neil",
"Nettie", "Piggies", "Psylocke", "Queen", "Ramble", "Sinv-25",
"Sinv12", "Sol-11", "Sol-18", "Sol-20", "Sol-49", "Sol-6", "Sol-J1",
"Sol-M2", "Sol-M3", "SolB8", "st_stephen", "Starr", "Sun_King",
"Taxman", "Tombstone_", "Wallflower", "Weight", "Wigwam", "Workingman",
"Yellow"), class = "factor"), All = c(0.3357, 0.4166, 0.0242,
0.9708, 0.4518, 0.0666, 0, 0.5925, 0.2349, 0.3242, 0.3278, 0.0246,
0.0352, 0.0646, 0.0563, 0.6854, 0.4664, 0.8298, 0.8831, NA, 0.0078,
0.771, 0.1376, 0.0055, 2e-04, 1, 0.2577, 0.3326, 0.0066, 0.0024,
0.6136, 0.4155, 0.9931, 0.2111, 0.6373, 0.0762, 0.5153, 0.1103,
0.0569, 0, 1, 0.732, 0, 0, 0.9225, 0.1257, 0.1658, 0.9603, 0.4629,
0, 0.4155, 0.0791, 0.0777, 0.3996, 0.1212, 0.6207, 0.5766, 0,
0.1347, 0.3754, 0, 0.2737), X1_only = c(0.4758, 0.3188, 0.1465,
0.3209, 1, 0.0278, 0.2065, 0.6187, 0.9377, 0.8586, 0.7292, 0.0133,
1, 1, 1, 0.961, 0.427, 1, 0.2203, NA, 0.8919, 0.2695, 0.3724,
0.5798, 0.5304, 1, 0.568, 0.2291, 0.4376, 1, 0.1572, 0.2022,
0.9544, 0.2462, 0.9699, 0.4439, 0.2204, 0.1135, 0.3063, 0.0311,
0.0384, 0.2833, 1, 0.7024, 0.7382, 0.4923, 0.4453, 0.2341, 0.7493,
0.0868, 0.8801, 0.8708, 1, 1, 1, 0.0491, 0.291, 0.2037, 0.1342,
0.5321, 0.4787, 0.7801), X78_only = c(0.3379, 0.4102, 0.2134,
0.6807, 0.8242, 1, 0.0046, 0.279, 0.825, 0.7563, 0.6055, 0.7472,
1, 0.4958, 0.0018, 0.0175, 1, 1, 0.5647, NA, 0.2124, 0.519, 0.5204,
0.2272, 0.03, 1, 0.0319, NA, 0.4467, 0.4473, 0.1593, 0.6066,
0.5907, 0.0624, 0.5699, 0.6585, 0.1414, 0.546, 0.6395, 0.0102,
0.3112, 0.791, 0, 0.7753, 0.4155, 0.9279, 0.4834, 0.3059, 0.5967,
0.373, 0.4114, 0.9291, 0.1159, 0.7238, 0.5993, 0.7975, 0.3283,
0.0511, 0.4902, 0.0438, 2e-04, 0.2357), X8_removed = c(0.0967,
0.5831, 0.058, 0.9268, 0.3518, 0.0629, 0, 0.6229, 0.2217, 0.2602,
0.7123, 0.0181, 0.0348, 0.1482, 0.1706, 0.6748, 0.3238, 0.8134,
0.8032, NA, 0.0246, 0.5794, 0.5204, 0.0254, 0.0056, 1, 0.6597,
0.3373, 0.004, 0.0087, 0.9061, 0.577, 0.9565, 0.4168, 0.7951,
0.1069, 0.4071, 0.1457, 0.1453, 0, 0.8385, 0.4658, 0, 0, 0.7396,
0.0748, 0.3677, 0.9571, 0.1188, 0, 0.5673, 0.0396, 0.0708, 0.3645,
0.1147, 0.5851, 1, 0.001, 0.0614, 0.131, 0, 0.4813), X8_only = c(0.1169,
0.8327, 0.2169, 0.0907, 1, 1, 0.07, 0.486, 0.709, 0.8882, 0.4389,
1, 0.7078, 0.4496, 0.1266, 0.1945, 0.4527, 1, 0.6518, NA, 0.3594,
0.7715, 0.134, 0.2389, 0.0203, 1, 0.1061, NA, 0.1293, 0.2558,
0.167, 0.4815, 0.7756, 0.0403, 0.2448, 0.2265, 0.0952, 0.6658,
0.3405, 0.0402, 0.5906, 0.2405, 0.0086, 0.5086, 0.4709, 1, 0.0567,
0.4146, 0.7554, 0.104, 0.1917, 0.8625, 1, 1, 1, 0.8727, 0.1439,
0.0452, 1, 0.5804, 0, 0.2764), X7_removed = c(0.2989, 0.7268,
0.0087, 0.8874, 0.5853, 0.0568, 0, 0.7622, 0.4226, 0.3232, 0.3972,
0.02, 0.0159, 0.0541, 0.4919, 0.5951, 0.5525, 0.8114, 0.5738,
NA, 0.0062, 0.7274, 0.0155, 0.0233, 0.002, 1, 0.232, 0.3476,
0.011, 9e-04, 0.5433, 0.3725, 0.7263, 0.2462, 0.4556, 0.0426,
0.7468, 0.1235, 0.0051, 0, 1, 0.8962, 0.0014, 0, 0.9892, 0.1163,
0.1284, 0.6873, 0.3932, 0, 0.3722, 0.0889, 0.3782, 0.4761, 0.0484,
0.5321, 0.5519, 0, 0.3453, 0.0732, 0, 0.3483), X7_only = c(1,
0.5714, 0.2825, 0.8673, 0.5557, 0.6861, 0.0044, 0.1146, 0.4957,
0.5248, 0.8372, 0.6665, 0.6789, 1, 0.0082, 0.1759, 0.3719, 1,
0.704, NA, 0.2585, 0.4634, 0.4283, 0.6815, 0.4161, 1, 0.1691,
NA, 0.4563, 1, 0.226, 1, 0.2349, 0.5886, 0.8154, 0.8839, 0.1631,
1, 0.5112, 0.1529, 1, 0.7245, 4e-04, 0.3095, 0.6184, 0.5542,
0.749, 0.394, 0.0298, 0.1994, 0.2881, 0.7696, 0.0637, 0.652,
1, 0.1494, 1, 0.3283, 0.134, 0.1992, 0.0848, 0.5826), X5_removed = c(1,
0.1453, 0.0176, 0.8428, 0.2277, 0.2563, 0, 0.5326, 0.1549, 0.4405,
0.395, 0.0195, 0.08, 0.1069, 0.0316, 0.6298, 0.5157, 1, 0.5967,
NA, 0.0265, 0.5703, 0.2667, 0.3485, 0.0021, 1, 0.1821, 0.3006,
0.007, 0.0112, 0.1964, 0.4427, 0.769, 0.1214, 0.6064, 0.0914,
0.4188, 0.021, 0.0814, 0, 0.8372, 0.8052, 0, 0, 0.8662, 0.7917,
0.0924, 0.9316, 0.7399, 0, 0.2031, 0.0701, 0.0652, 0.6636, 0.0513,
0.2049, 0.7161, 0, 0.0407, 0.1729, 0, 0.3079), X5_only = c(0.0642,
0.631, 0.5193, 0.979, 0.5348, 0.1304, 0.02, 0.0217, 0.0871, 0.2022,
0.7602, 1, 0.3532, 0.5292, 1, 0.3677, 0.0896, 0.3702, 0.6084,
NA, 0.1518, 0.3467, 0.1171, 0.0252, 0.7894, 1, 0.9842, 0.7315,
0.8511, 0.0717, 0.0585, 0.7955, 0.3517, 1, 0.8263, 0.6102, 0.268,
0.1071, 0.3837, 0.0175, 0.5887, 1, NA, 0.1198, 0.8537, 0.0101,
0.3807, 0.4939, 0.1469, 0.1368, 0.5458, 0.2514, 1, 0.3692, 1,
0.4877, 0.5787, 0.6025, 0.5888, 1, 0.3472, 1), X4_removed = c(0.4492,
0.3821, 0.0121, 0.9957, 0.5158, 0.0498, 0, 0.718, 0.8003, 0.1716,
0.661, 0.0194, 0.0511, 0.1862, 0.0188, 0.6454, 0.5077, 1, 0.8794,
NA, 0.3458, 0.6059, 0.1315, 0.0099, 0.003, 1, 0.0585, 0.4635,
0.0357, 0.0289, 0.6835, 0.2247, 0.8437, 0.3585, 0.6074, 0.1926,
0.3432, 0.3615, 0.0322, 0, 0.8418, 0.7076, 0, 0.9281, 0.7697,
0.1011, 0.3068, 0.971, 0.4686, 0, 0.3731, 0.1024, 0.0683, 0.8112,
0.3742, 0.7381, 0.2738, 0.0089, 0.2366, 0.6924, 0, 0.1984), X4_only = c(0.6485,
0.0709, 0.1639, 0.6908, 1, 1, 0.4469, 0.639, 0.0378, 0.5116,
0.0026, 0.6549, 0.6928, 0.2884, 1, 0.4386, 0.6246, 0.6188, 1,
NA, 0.0966, 0.3946, 0.7223, 0.1357, 0.8912, 1, 0.4741, 0.7526,
0.2005, 0.013, 1, 0.455, 0.1086, 0.1184, 0.8975, 0.3181, 0.9958,
0.0644, 0.0975, 0.0721, 1, 1, 1, 7e-04, 0.2754, 0.4852, 0.065,
0.747, 0.4823, 0.1971, 0.6178, 0.3781, 1, 0.362, 0.1168, 0.382,
0.4267, 8e-04, 0.188, 0.2115, 0.2937, 1), X3_removed = c(0.3009,
0.3414, 0.02, 0.9935, 0.4216, 0.1273, 0, 0.6406, 0.2728, 0.5307,
0.477, 0.0612, 0.0627, 0.0808, 0.1636, 0.6506, 0.6507, 0.8122,
0.9531, NA, 0.0144, 0.9274, 0.1646, 0.0171, 1e-04, 1, 0.2732,
0.4153, 0.0141, 0.0105, 0.6892, 0.3701, 0.9956, 0.0418, 0.5436,
0.2755, 0.4803, 0.0959, 0.1199, 0, 0.833, 0.5373, 0, 0, 0.9701,
0.1054, 0.1558, 0.9964, 0.6849, 0, 0.2023, 0.1072, 0.3401, 0.3629,
0.2504, 0.6056, 0.5372, 2e-04, 0.1168, 1, 0, 0.242), X3_only = c(1,
0.9325, 0.772, 0.5505, 1, 0.2068, 0.0829, 0.17, 0.8951, 0.0225,
0.8263, 0.2111, 0.5087, 0.768, 0.2471, 0.6294, 0.2815, 1, 0.0496,
NA, 0.3364, 0.6286, 0.2102, 0.6816, 0.372, 1, 0.7311, 0.5138,
0.0683, 0.1996, 0.6998, 1, 0.6988, 0.4426, 0.6669, 0.0412, 0.6081,
1, 0.237, 6e-04, 0.6349, 0.7124, 1, 0.2314, 0.0398, 1, 0.3487,
0.8153, 0.1271, 0.1145, 0.8641, 0.4056, 0.1488, 1, 0.2357, 0.26,
1, 0.2678, 0.5537, 0.0317, 0.0467, 1), X2_removed = c(0.6335,
0.349, 0.2095, 0.9777, 0.8928, 0.0571, 0, 0.4285, 0.2036, 0.3168,
0.3668, 0.0854, 0.413, 0.0608, 0.0526, 0.7608, 0.3094, 0.8186,
0.9273, NA, 0.0014, 0.6512, 0.4424, 0.0275, 0.2121, 1, 0.3008,
0.2381, 0.0173, 0.0075, 0.7423, 0.6126, 0.979, 0.1716, 0.862,
0.0245, 0.5096, 0.2795, 0.4794, 0, 1, 0.6888, 0, 0, 0.6213, 0.0935,
0.1351, 0.6946, 0.4708, 0.1458, 0.899, 0.4391, 0.0727, 0.5004,
0.3974, 0.8854, 0.2696, 0, 0.1846, 0.5871, 0, 0.2966), X2_only = c(0.191,
0.4397, 0.0403, 0.3606, 0.0089, 1, 0.0033, 0.659, 0.1818, 0.0949,
0.5521, 0.1637, 0.0014, 1, NA, 0.8585, 1, 1, 0.9437, NA, 0.4086,
0.1699, 0.0648, 0.9087, 0.0011, 1, 0.1291, 0.5329, 0.2315, 0.2844,
0.6429, 0.0488, 0.1814, 0.8658, 0.0869, 0.8394, 0.5938, 0.1722,
0, 0.0098, 1, 1, 1, 0.1742, 0.3911, 0.8523, 0.7331, 0.1271, 0.5119,
0, 0.0105, 0.0035, 1, 0.5665, 0.072, 0.2928, 0.4224, 0.5491,
0.4274, 0.1054, 0, 0.5817), X1_removed = c(0.1653, 0.7658, 0.0718,
0.7705, 0.4193, 0.1894, 0, 0.5167, 0.1053, 0.2823, 0.0496, 0.1439,
0.0258, 0.0676, 0.031, 0.5465, 0.4909, 0.6464, 0.9383, NA, 0.0124,
0.9288, 0.069, 0.0116, 6e-04, 1, 0.3301, 0.508, 0.0175, 8e-04,
0.6016, 0.7442, 0.9609, 0.4151, 0.6049, 0.1266, 0.4281, 0.2719,
0.0039, 0, 0.315, 1, 0, 0, 0.8931, 0.1124, 0.3804, 0.9233, 0.3355,
0, 0.3542, 0.0363, 0.0679, 0.2652, 0.122, 0.4025, 0.8155, 2e-04,
0.2642, 0.3629, 0, 0.2897)), .Names = c("Loci", "All", "X1_only",
"X78_only", "X8_removed", "X8_only", "X7_removed", "X7_only",
"X5_removed", "X5_only", "X4_removed", "X4_only", "X3_removed",
"X3_only", "X2_removed", "X2_only", "X1_removed"), class = "data.frame", row.names = c(NA,
-62L))
#now make subsets of this big dataset
split1_data<-droplevels(subset(data,data$Loci %in% data$Loci[1:8]))
split2_data<-droplevels(subset(data,data$Loci %in% data$Loci[9:16]))
split3_data<-droplevels(subset(data,data$Loci %in% data$Loci[17:24]))
split4_data<-droplevels(subset(data,data$Loci %in% data$Loci[25:32]))
split5_data<-droplevels(subset(data,data$Loci %in% data$Loci[33:40]))
split6_data<-droplevels(subset(data,data$Loci %in% data$Loci[41:48]))
split7_data<-droplevels(subset(data,data$Loci %in% data$Loci[49:56]))
split8_data<-droplevels(subset(data,data$Loci %in% data$Loci[57:62]))
#and melt each of them
split1_datam<-melt(split1_data,id="Loci")
split2_datam<-melt(split2_data,id="Loci")
split3_datam<-melt(split3_data,id="Loci")
split4_datam<-melt(split4_data,id="Loci")
split5_datam<-melt(split5_data,id="Loci")
split6_datam<-melt(split6_data,id="Loci")
split7_datam<-melt(split7_data,id="Loci")
split8_datam<-melt(split8_data,id="Loci")
#and make a plot for each of the melted subsets
p1<- ggplot(split1_datam, aes(x =Loci, y = value, color = variable, width=.15))+ geom_bar(position="dodge")+ geom_hline(yintercept=0.05)+ opts(legend.position="none",axis.text.x = theme_text(angle=90, size=8)) + scale_y_discrete(breaks=seq(0,1)) + ylab(NULL)
p2<- ggplot(split2_datam, aes(x =Loci, y = value, color = variable, width=.15)) + geom_bar(position="dodge") + geom_hline(yintercept=0.05)+ opts(legend.position = "none", axis.text.x = theme_text(angle=90, size=8)) + scale_y_discrete(breaks=seq(0,1))+ scale_fill_grey() + ylab(NULL)
p3<-p <- ggplot(split3_datam, aes(x =Loci, y = value, color = variable, width=.15))
p3<-p3 + geom_bar(position="dodge") + geom_hline(yintercept=0.05)+ opts(legend.position = "none", axis.text.x = theme_text(angle=90, size=8)) + scale_y_discrete(breaks=seq(0,1)) + ylab(NULL)
p4<-p <- ggplot(split4_datam, aes(x =Loci, y = value, color = variable, width=.15))
p4<-p4 + geom_bar(position="dodge") + geom_hline(yintercept=0.05)+ scale_y_discrete(breaks=seq(0,1))+opts(legend.position="none", axis.text.x = theme_text(angle=90, size=8)) + ylab(NULL)
p5<-p <- ggplot(split5_datam, aes(x =Loci, y = value, color = variable, width=.15))
p5<-p5 + geom_bar(position="dodge") + geom_hline(yintercept=0.05)+ opts(legend.position = "none", axis.text.x = theme_text(angle=90, size=8))+ scale_y_discrete(breaks=seq(0,1)) + ylab(NULL)
p6<-p <- ggplot(split6_datam, aes(x =Loci, y = value, color = variable, width=.15))
p6<-p6 + geom_bar(position="dodge") + geom_hline(yintercept=0.05) + scale_y_discrete(breaks=seq(0,1))+ opts(legend.position = "none", axis.text.x = theme_text(angle=90, size=8)) + ylab(NULL)
p7<-p <- ggplot(split7_datam, aes(x =Loci, y = value, color = variable, width=.15))
p7<-p7 + geom_bar(position="dodge") + geom_hline(yintercept=0.05) + scale_y_discrete(breaks=seq(0,1))+ opts(legend.position = "none", axis.text.x = theme_text(angle=90, size=8)) + ylab(NULL)
p8<-p <- ggplot(split8_datam, aes(x =Loci, y = value, color = variable, width=.15))
p8<-p8 + geom_bar(position="dodge") + geom_hline(yintercept=0.05) + scale_y_discrete(breaks=seq(0,1))+opts(legend.position="none",axis.text.x = theme_text(angle=90, size=8)) + ylab(NULL)
#make a bad attempt at creating a legend for the entire multiplot?
leg <- ggplotGrob(p1 + opts(keep="legend_box"))
## one needs to provide the legend with a well-defined width
legend=gTree(children=gList(leg), cl="legendGrob")
widthDetails.legendGrob <- function(x) unit(2, "cm")
#plot the first four plots together
pdf("sensivitity_page_2_plots.pdf")
grid.arrange(p1, p2, p3, p4, nrow=2, legend=legend, main ="Sensitivity", left = "Pvalue")
dev.off()
#same with the next four
pdf("sensivitity_page_2_plots.pdf")
leg <- ggplotGrob(p5 + opts(keep="legend_box"))
## one needs to provide the legend with a well-defined width
legend=gTree(children=gList(leg), cl="legendGrob")
widthDetails.legendGrob <- function(x) unit(2, "cm")
grid.arrange(p5, p6, p7, p8, nrow=2, legend=legend, main ="Sensitivity", left = "Pvalue")
dev.off()
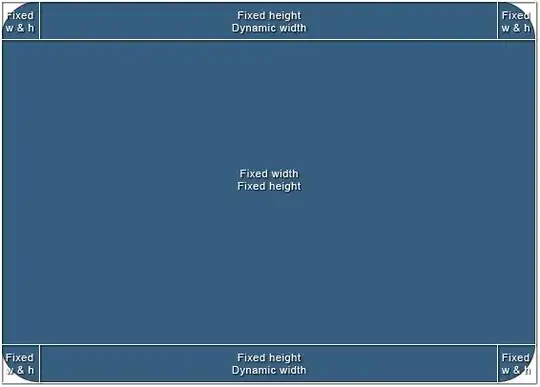
The plots will look like this:
I have the following questions:
- How can I space the y-axis label so that it's not so close to the edge?
- How can I do the same for the main title
- Why is the legend not appearing? (that portion of the code was lifted from the grid.arrange documentation). And, when it does, how do I space that properly? Anybody else out there use grid.arrange?
- If not, is there another way to apply layout()-like positioning in ggplot?
Thanks so much, everyone!