I am not sure how to do the prediction for some new data using trained Gaussian Mixture Model (GMM). For example, I have got some labelled data drawn from 3 different classes (clusters). For each class of data points, I fit a GMM (gm1, gm2 and gm3). Suppose we know the number of Gaussian mixture for each class (e.g., k1=2, k2=1 and k3=3) or it can be estimated (optimised) using Akaike information criterion (AIC). Then when I have got some new dataset, how can I know if it is more likely belong to class 1, 2 or 3?
Some Matlab script shows what I mean:
clc; clf; clear all; close all;
%% Create some artificial training data
% 1. Cluster 1 with two mixture of Gaussian (k1 = 2)
rng default; % For reproducibility
mu1 = [1 2];
sigma1 = [3 .2; .2 2];
mu2 = [-1 -2];
sigma2 = [2 0; 0 1];
X1 = [mvnrnd(mu1,sigma1,200); mvnrnd(mu2,sigma2,100)];
options1 = statset('Display', 'final');
k1 = 2;
gm1 = fitgmdist(X1, k1, 'Options', options1);
% 2. Cluster 2 with one mixture of Gaussian (k2 = 1)
mu3 = [6 4];
sigma3 = [3 .1; .1 4];
X2 = mvnrnd(mu3,sigma3,300);
options2 = statset('Display', 'final');
k2 = 1;
gm2 = fitgmdist(X2, k2, 'Options', options2);
% 3. Cluster 3 with three mixture of Gaussian (k3 = 3)
mu4 = [-5 -6];
sigma4 = [1 .1; .1 1];
mu5 = [-5 -10];
sigma5 = [6 .1; .1 1];
mu6 = [-2 -15];
sigma6 = [8 .1; .1 4];
X3 = [mvnrnd(mu4,sigma4,200); mvnrnd(mu5,sigma5,300); mvnrnd(mu6,sigma6,100)];
options3 = statset('Display', 'final');
k3 = 3;
gm3 = fitgmdist(X3, k3, 'Options', options3);
% Display
figure,
scatter(X1(:,1),X1(:,2),10,'ko'); hold on;
ezcontour(@(x,y)pdf(gm1, [x y]), [-12 12], [-12 12]);
scatter(X2(:,1),X2(:,2),10,'ko');
ezcontour(@(x,y)pdf(gm2, [x y]), [-12 12], [-12 12]);
scatter(X3(:,1),X3(:,2),10,'ko');
ezcontour(@(x,y)pdf(gm3, [x y]), [-12 12], [-12 12]); hold off;
We can get the figure:
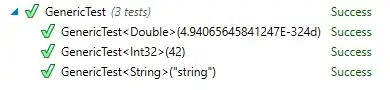
Then we have got some new testing data for example:
%% Create some artificial testing data
mut1 = [6.1 3.8];
sigmat1 = [3.1 .1; .1 4.2];
mut2 = [5.8 4.5];
sigmat2 = [2.8 .1; .1 3.8];
Xt1 = [mvnrnd(mut1,sigmat1,500); mvnrnd(mut2,sigmat2,100)];
figure,
scatter(Xt1(:,1),Xt1(:,2),10,'ko');
xlim([-12 12]); ylim([-12 12]);

I made the testing data similar to the Cluster 2 data on purpose. After we do the training using GMM, can we somehow predict the label of the new testing data? Is that possible to get some probabilities out like (p1 = 18%, p2 = 80% and p3 = 2%) for the prediction of each class. As we have got p2=80%, we can then have a hard classification that the new testing data is labelled as Cluster 2.
p.s.: I have found this post but it seems to theoretical to me (A similar post). If you can please put some simple Matlab script in your reply.
Thanks very much. A.
EDIT:
As Amro replied a solution for the problem, I have got more questions.
Amro created a new GMM using the entire dataset with some initialisation:
% initial parameters of the new GMM (combination of the previous three) % (note PComponents is normalized according to proportion of data in each subset) S = struct('mu',[gm1.mu; gm2.mu; gm3.mu], ... 'Sigma',cat(3, gm1.Sigma, gm2.Sigma, gm3.Sigma), ... 'PComponents',[gm1.PComponents*n1, gm2.PComponents*n2, gm3.PComponents*n3]./n); % train the final model over all instances opts = statset('MaxIter',1000, 'Display','final'); gmm = fitgmdist(X, k, 'Options',opts, 'Start',S);What Amro got is something like below
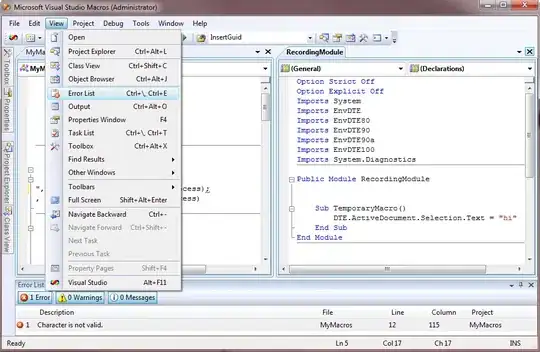
which may not be suitable of my data because it separates my labelled cluster1, and cluster2 mixed with part of cluster1. This is what I am trying to avoid.
Here what I present is an artificial numerical example; however, in my real application, it deals with problem of image segmentation (for example, cluster1 is my background image and cluster2 is the object I want to separate). Then I try to somehow 'force' the separate GMM to fit separate classes. If two clusters are far away (for example, cluster1 and cluster 3 in this example), there is no problem to use Amro's method to combine all the data and then do a GMM fitting. However, when we do the training on the image data, it will never be perfect to separate background from object due to the limitation of the resolution (caused partial volume effect); therefore, it is very likely we have the case of cluster1 overlapped with cluster2 as shown. I think maybe mix all the data and then do the fitting will cause some problem for further prediction of the new data, am I right?
However, after a little bit of thinking, what I am trying to do now is:
% Combine the mixture of Gaussian and form a new gmdistribution muAll = [gm1.mu; gm2.mu; gm3.mu]; sigmaAll = cat(3, gm1.Sigma, gm2.Sigma, gm3.Sigma); gmAll = gmdistribution(muAll, sigmaAll); pt1 = posterior(gmAll, Xt1);What do you guys think? Or it is equivalent to Amro's method? If so, is there a method to force my trained GMM separated?
Also, I have question about the rationale of using the
posteriorfunction. Essentially, I want to estimate the likelihood of my testing data given the GMM fitting. Then why we calculate the posterior probability now? Or it is just a naming issue (in other words, the 'posterior probability'='likelihood')?As far as I know, GMM has always been used as a unsupervised method. Someone even mentioned to me that GMM is a probability version of k-means clustering. Is it eligible to use it in such a 'supervised' style? Any recommended papers or references?
Thanks very much again for your reply! A.