Note: this more of an extended comment than an answer.
You encountered a problem I also just encountered whilst developing a NEAT version for javascript. The original paper published in ~2002 is very unclear.
The original paper contains the following:
Whenever a new
gene appears (through structural mutation), a global innovation number is incremented
and assigned to that gene. The innovation numbers thus represent a chronology of the
appearance of every gene in the system. [..] ; innovation numbers are never changed. Thus, the historical origin of every
gene in the system is known throughout evolution.
But the paper is very unclear about the following case, say we have two ; 'identical' (same structure) networks:
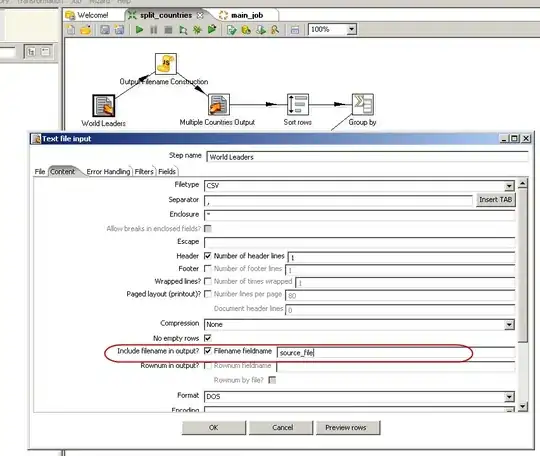
The networks above were initial networks; the networks have the same innovation ID, namely [0, 1]. So now the networks randomly mutate an extra connection.

Boom! By chance, they mutated to the same new structure. However, the connection ID's are completely different, namely [0, 2, 3] for parent1 and [0, 4, 5] for parent2 as the ID is globally counted.
But the NEAT algorithm fails to determine that these structures are the same. When one of the parents scores higher than the other, it's not a problem. But when the parents have the same fitness, we have a problem.
Because the paper states:
In composing the offspring, genes are randomly chosen from veither parent at matching genes, whereas all excess or disjoint genes are always included from the more fit parent, or if they are equally fit, from both parents.
So if the parents are equally fit, the offspring will have connections [0, 2, 3, 4, 5]. Which means that some nodes have double connections... Removing global innovation counters, and just assign id's by looking at node_in and node_out, you avoid this problem.
So when you have equally fit parents, yes you have optimized the algorithm. But this is almost never the case.
Quite interesting: in the newer version of the paper, they actually removed that bolded line! Older version here.
By the way, you can solve this problem by instead of assigning innovation ID's, assign ID based on node_in and node_out using pairing functions. This creates quite interesting neural networks when fitness is equal: