I'm trying to create dendrograms from two different distance matrices and compare them. I used the code here as a starting point, but the problem is since I'm using two different matrices but same clustering method, I need to plot two different matrices together for a comparative analysis. I was wondering if it is possible to separate to halves of each square/node diagonally to show two different distance matrices.
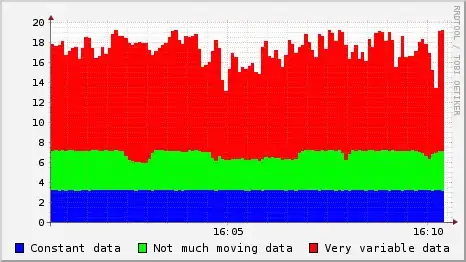
This image represents the result which I'm targeting for:
Here is my code:
from sklearn import preprocessing
from sklearn.neighbors import DistanceMetric
import pandas as pd
import numpy as np
from ete3 import Tree
from sklearn.metrics.pairwise import cosine_similarity
from sklearn.metrics.pairwise import cosine_distances
import scipy
import pylab
import scipy.cluster.hierarchy as sch
import scipy.spatial.distance as sd
import random
#g[n] is a one dimensional array containing datapoints
g1 = random.sample(range(30), 5)
g2 = random.sample(range(30), 5)
g3 = random.sample(range(30), 5)
g4 = random.sample(range(30), 5)
g5 = random.sample(range(30), 5)
g1 = np.array(g1)
g2 = np.array(g2)
g3 = np.array(g3)
g4 = np.array(g4)
g5 = np.array(g5)
X = (g1,g2,g3,g4,g5)
#Comparing between euclidean and cosine###########################################
distanceC = cosine_distances(X)
dist = DistanceMetric.get_metric('euclidean')
distanceE = dist.pairwise(X)
##################################################################################
#Plots############################################################################
# Compute and plot first dendrogram.
fig = pylab.figure(figsize=(8,8))
ax1 = fig.add_axes([0.09,0.1,0.2,0.6])
Y = sch.average(sd.squareform(distanceC))
Z1 = sch.dendrogram(Y, orientation='right')
ax1.set_xticks([])
ax1.set_yticks([])
# Compute and plot second dendrogram.
ax2 = fig.add_axes([0.3,0.71,0.6,0.2])
Y = sch.average(sd.squareform(distanceE))
Z2 = sch.dendrogram(Y)
ax2.set_xticks([])
ax2.set_yticks([])
# Plot distance matrix.
axmatrix = fig.add_axes([0.3,0.1,0.6,0.6])
idx1 = Z1['leaves']
idx2 = Z2['leaves']
distance = distance[idx1,:]
distance = distance[:,idx2]
im = axmatrix.matshow(distance, aspect='auto', origin='lower', cmap=pylab.cm.YlGnBu)
axmatrix.set_xticks([])
axmatrix.set_yticks([])
# Plot colorbar.
axcolor = fig.add_axes([0.91,0.1,0.02,0.6])
pylab.colorbar(im, cax=axcolor)
fig.show()
fig.savefig('dendrogram.png')
##################################################################################