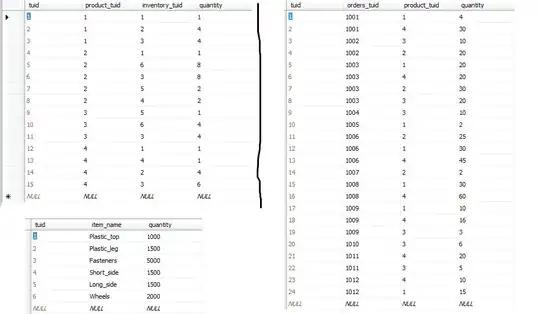
I found a way to convert igraph into visNetwork (refer to Interactive arules with arulesViz and visNetwork). Suppose before and after conversion from igraph to visNetwork should be the same, but my result shows after convert into visNetwork, the results are different.
I'll try demonstrate the issue using sample data data("Groceries") from Library(arules).
#Pre-defined library
library(arules)
library(arulesViz)
library(visNetwork)
library(igraph)
#Get sample data & get association rules
data("Groceries")
rules <- apriori(Groceries, parameter=list(support=0.01, confidence=0.4))
rules <- head(sort(rules, by="lift"), 10)
#Convert rules to data.table
library(data.table)
rules_dt <- data.table( lhs = labels( lhs(rules) ),
rhs = labels( rhs(rules) ),
quality(rules) )[ order(-lift), ]
print all rules in table format (sort by lift)
Plot top 10 association rules via using igraph
ig <- plot(rules, method="graph", control=list(type="items"))
Note: Based on association rules i plot a network diagram via using igraph, everything is correct. Next i'll try to convert existing igraph to visNetwork, then we compare the results.
tf <- tempfile( )
saveAsGraph(rules, file = tf, format = "dot" )
# clean up temp file if desired
#unlink(tf)
# Convert igraph to dataframe
ig_df <- as_data_frame(ig, what = "both")
# Plot visNetwork
visNetwork(
nodes = data.frame(
id = ig_df$vertices$name
,value = ig_df$vertices$lift # could change to lift or confidence
,title = ifelse(ig_df$vertices$label == "",ig_df$vertices$name,
ig_df$vertices$label)
,ig_df$vertices
),
edges = ig_df$edges
) %>%
visEdges(arrows ="to") %>%
visOptions( highlightNearest = T )
Plot top 10 association rules via using visNetwork

Note: For visNetwork diagram, the size of intercept node indicate "lift", the higher the lift, the larger the size of intercept node; unlike igraph diagram, size of intercept node indicate "support", while colour of intercept node indicate "lift".
Let's compare the igraph and visNetwork
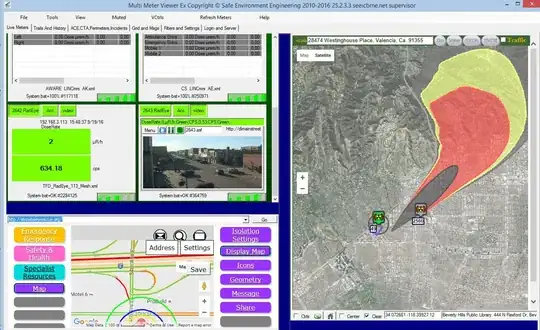 By referring to the association rules in table format, rules no.10 (rules with smallest "lift"), suppose the size of the intercept node is the smallest, but end up it's not the smallest.
By referring to the association rules in table format, rules no.10 (rules with smallest "lift"), suppose the size of the intercept node is the smallest, but end up it's not the smallest.
Problems
I tried to further drill down into ig_df <- get.data.frame( ig, what = "both" ), and i found something weird on ig_df$vertices table, which generated from as_data_frame function from library(igraph).
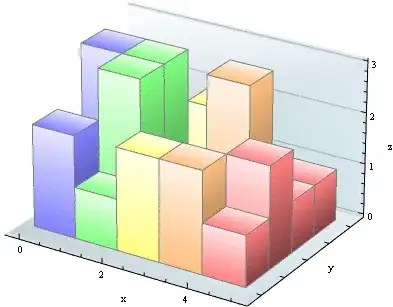 I found that the assoc10 (intercept node for association rules no.10) actually had
I found that the assoc10 (intercept node for association rules no.10) actually had NA for all variables (i.e. "support", "confidence", "lift" & "count"), more precisely the dimension for columns "support", "confidence", "lift" & "count" in ig_df$vertices are moving up one row! Kindly correct me if i'm wrong..
Conclusion
Since the key to convert an igraph into visNetwork is to use this as_data_frame to get extract all data from an igraph and convert those data into dataframe, then plot visNetwork using the data from extracted dataframe. But due to extract issue when using as_data_frame to extract data from igraph, so the result from also different.
Question: is this a bug? or i made a mistake on my code? Any suggestion is welcome. Thanks!