I have following set of points that lie on a boundary and want to create the polygon that connects these points. For a person it is quite obvious what path to follow, but I am unable to find an algorithm that does the same and trying to solve it myself it all seems quite tricky and ambiguous occasionally. What is the best solution for this?
As a background.
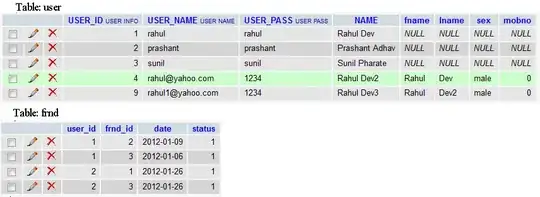
This is the boundary for the julia set with constant = -0.624+0.435j with stable area defined after 100 iterations. I got these points by setting the stable points to 1 and all other to zero and then convolving with a 3x3 matrix [[1, 1, 1], [1, 1, 1], [1, 1, 1]] and select the points that have value 1. My experimenting code is as follows:
import numpy as np
from scipy.signal import convolve2d
import matplotlib.pyplot as plt
r_min, r_max = -1.5, 1.5
c_min, c_max = -2.0, 2.0
dpu = 50 # dots per unit - 50 dots per 1 units means 200 points per 4 units
max_iterations = 100
cmap='hot'
intval = 1 / dpu
r_range = np.arange(r_min, r_max + intval, intval)
c_range = np.arange(c_min, c_max + intval, intval)
constant = -0.624+0.435j
def z_func(point, constant):
z = point
stable = True
num_iterations = 1
while stable and num_iterations < max_iterations:
z = z**2 + constant
if abs(z) > max(abs(constant), 2):
stable = False
return (stable, num_iterations)
num_iterations += 1
return (stable, 0)
points = np.array([])
colors = np.array([])
stables = np.array([], dtype='bool')
progress = 0
for imag in c_range:
for real in r_range:
point = complex(real, imag)
points = np.append(points, point)
stable, color = z_func(point, constant)
stables = np.append(stables, stable)
colors = np.append(colors, color)
print(f'{100*progress/len(c_range)/len(r_range):3.2f}% completed\r', end='')
progress += len(r_range)
print(' \r', end='')
rows = len(r_range)
start = len(colors)
orig_field = []
for i_num in range(len(c_range)):
start -= rows
real_vals = [color for color in colors[start:start+rows]]
orig_field.append(real_vals)
orig_field = np.array(orig_field, dtype='int')
rows = len(r_range)
start = len(stables)
stable_field = []
for i_num in range(len(c_range)):
start -= rows
real_vals = [1 if val == True else 0 for val in stables[start:start+rows]]
stable_field.append(real_vals)
stable_field = np.array(stable_field, dtype='int')
kernel = np.array([[1, 1, 1], [1, 1, 1], [1, 1, 1]])
stable_boundary = convolve2d(stable_field, kernel, mode='same')
boundary_points = []
cols, rows = stable_boundary.shape
assert cols == len(c_range), "check c_range and cols"
assert rows == len(r_range), "check r_range and rows"
zero_field = np.zeros((cols, rows))
for col in range(cols):
for row in range(rows):
if stable_boundary[col, row] in [1]:
real_val = r_range[row]
# invert cols as min imag value is highest col and vice versa
imag_val = c_range[cols-1 - col]
stable_boundary[col, row] = 1
boundary_points.append((real_val, imag_val))
else:
stable_boundary[col, row] = 0
fig, ((ax1, ax2), (ax3, ax4)) = plt.subplots(nrows=2, ncols=2, figsize=(5, 5))
ax1.matshow(orig_field, cmap=cmap)
ax2.matshow(stable_field, cmap=cmap)
ax3.matshow(stable_boundary, cmap=cmap)
x = [point[0] for point in boundary_points]
y = [point[1] for point in boundary_points]
ax4.plot(x, y, 'o', c='r', markersize=0.5)
ax4.set_aspect(1)
plt.show()
Output with dpu = 200 and max_iterations = 100:
inspired by this Youtube video: What's so special about the Mandelbrot Set? - Numberphile