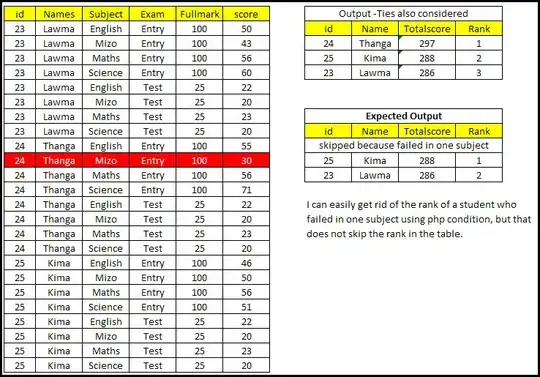
Is there a way to get from a to b in the following figure with scripting? I am using seaborn.clustermap() to get to a (i.e. the order of the rows are preserved. However, columns order change only at second highest level).
I was wondering whether it is possible to use the seaborn.matrix.ClusterGrid that is returned by seaborn.clustermap(), modify it and plot the modified results.
 b
P.S. The reason I am asking this is that the order has a meaning (first comes blue, next green, and finally red).
b
P.S. The reason I am asking this is that the order has a meaning (first comes blue, next green, and finally red).
Update: Here is a small data set to generate the situation:
df = pd.DataFrame([[1, 1.1, 0.9, 1.9, 2, 2.1, 2.8, 3, 3.1],
[1.8, 2, 2.1, 0.7, 1, 1.1, 2.7, 3, 3.3]],
columns = ['d1', 'd2', 'd3',
'l3', 'l2', 'l1',
'b1', 'b2', 'b3'],
index = ['p1', 'p2'])
cg = sns.clustermap(df); ## returns a ClusterGrid
The output is this:
We can think of columns starting with b as breakfast, l as lunch and d as dinner. Now, the order, is breakfast -> dinner -> lunch. I want to get to breakfast -> lunch -> dinner.