I have a specific performance issue, that i wish to extend more generally if possible.
Context:
I've been playing around on google colab with a python code sample for a Q-Learning agent, which associate a state and an action to a value using a defaultdict:
self._qvalues = defaultdict(lambda: defaultdict(lambda: 0))
return self._qvalues[state][action]
Not an expert but my understanding is it returns the value or add and returns 0 if the key is not found.
i'm adapting part of this in R.
the problem is I don't how many state/values combinations I have, and technically i should not know how many states I guess.
At first I went the wrong way, with the rbind of data.frames and that was very slow.
I then replaced my R object with a data.frame(state, action, value = NA_real).
it works but it's still very slow. another problem is my data.frame object has the maximum size which might be problematic in the future.
then I chanded my data.frame to a data.table, which gave me worst performance, then I finally indexed it by (state, action).
qvalues <- data.table(qstate = rep(seq(nbstates), each = nbactions),
qaction = rep(seq(nbactions), times = nbstates),
qvalue = NA_real_,
stringsAsFactors = FALSE)
setkey(qvalues, "qstate", "qaction")
Problem:
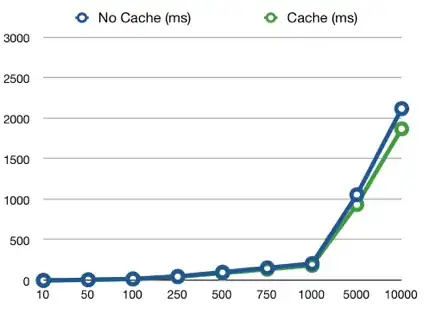
Comparing googlecolab/python vs my local R implementation, google performs 1000x10e4 access to the object in let's say 15s, while my code performs 100x100 access in 28s. I got 2s improvements by byte compiling but that's still too bad.
Using profvis, I see most of the time is spent accessing the data.table on these two calls:
qval <- self$qvalues[J(state, action), nomatch = NA_real_]$qvalue
self$qvalues[J(state, action)]$qvalue <- value
I don't really know what google has, but my desktop is a beast. Also I saw some benchmarks stating data.table was faster than pandas, so I suppose the problem lies in my choice of container.
Questions:
- is my use of a data.table wrong and can be fixed to improve and match the python implementation?
- is another design possible to avoid declaring all the state/actions combinations which could be a problem if the dimensions become too large?
- i've seen about the hash package, is it the way to go?
Thanks a lot for any pointer!
UPDATE:
thanks for all the input. So what I did was to replace 3 access to my data.table using your suggestions:
#self$qvalues[J(state, action)]$qvalue <- value
self$qvalues[J(state, action), qvalue := value]
#self$qvalues[J(state, action),]$qvalue <- 0
self$qvalues[J(state, action), qvalue := 0]
#qval <- self$qvalues[J(state, action), nomatch = NA_real_]$qvalue
qval <- self$qvalues[J(state, action), nomatch = NA_real_, qvalue]
this dropped the runtime from 33s to 21s
that's a massive improvement, but that's still extremely slow compared to the python defaultdict implementation.
I noted the following:
working in batch: I don't think I can do as the call to the function depends on the previous call.
peudospin> I see you are surprised the get is time consuming. so am I but that's what profvis states:
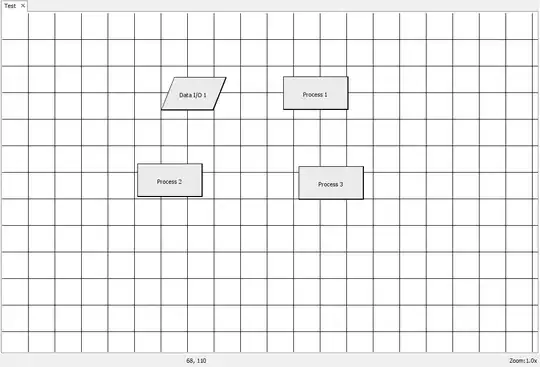 and here the code of the function as a reference:
and here the code of the function as a reference:
QAgent$set("public", "get_qvalue", function( state, action) {
#qval <- self$qvalues[J(state, action), nomatch = NA_real_]$qvalue
qval <- self$qvalues[J(state, action), nomatch = NA_real_, qvalue]
if (is.na(qval)) {
#self$qvalues[self$qvalues$qstate == state & self$qvalues$qaction == action,]$qvalue <- 0
#self$qvalues[J(state, action),]$qvalue <- 0
self$qvalues[J(state, action), qvalue := 0]
return(0)
}
return(qval)
})
At this point, if no more suggestion, I will conclude the data.table is just too slow for this kind of task, and I should look into using an env or a collections. (as suggested there: R fast single item lookup from list vs data.table vs hash )
CONCLUSION:
I replaced the data.table for a collections::dict and the bottleneck completely disappeared.