Edit: following interactions in the responses below, I believe there may be some issues with the plot() or plot.gam() functions when dealing with gam outputs. See responses below.
I am running a non parametric regression model <- gam(y ~ x, bs = "cs", data = data).
My data looks like what follows, where x is in logs. I have 273 observations
y x
[1,] 0.010234756 10.87952
[2,] 0.009165001 10.98407
[3,] 0.001330975 11.26850
[4,] 0.008000957 10.97803
[5,] 0.008579472 10.94924
[6,] 0.009746714 11.01823
I would like to plot the output of the model, basically the fitted curve. When I do
# graph
plot(model)
or
ggplot(data = data, mapping = aes(x = x y = y)) +
geom_point(size = 0.5, alpha = 0.5) +
geom_smooth(method="gam", formula= y~s(x, bs = "cs") )
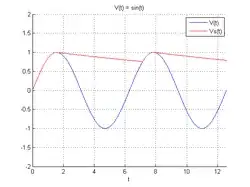
I get the desired output graphs (apologies for the original labels):
[
However, the two plotted curves are not exactly the same and I did not manage to find the parameters to tweak to remove the differences. Therefore I would like to plot the curve manually. Here it's my current attempt.
model <- gam(y~ s(x), bs = "cs", data = data)
names(model)
# summary(model)
model_fit <- as.data.frame(cbind(model$y, model$fitted.values,
model$linear.predictors, data$x,
model$residuals))
names(model_fit) <- c("y", "y_fit", "linear_pred", "x", "res")
### here the plotting
ggplot(model_fit) +
geom_point(aes(x = x, y = y_fit), size = 0.5, alpha = 0.5) +
geom_line(aes(x = x, y = y_fit))
However I get the following warning
geom_path: Each group consists of only one observation. Do you need to adjust the group aesthetic?
I do not seem to be able to fix the last graph (it seems the error is in geom_point() ) and add the confidence intervals, nor to find where to tweak the first two to make them exactly the same.