There are many ways to do this in R. Specifically, by, aggregate, split, and plyr, cast, tapply, data.table, dplyr, and so forth.
Broadly speaking, these problems are of the form split-apply-combine. Hadley Wickham has written a beautiful article that will give you deeper insight into the whole category of problems, and it is well worth reading. His plyr package implements the strategy for general data structures, and dplyr is a newer implementation performance tuned for data frames. They allow for solving problems of the same form but of even greater complexity than this one. They are well worth learning as a general tool for solving data manipulation problems.
Performance is an issue on very large datasets, and for that it is hard to beat solutions based on data.table. If you only deal with medium-sized datasets or smaller, however, taking the time to learn data.table is likely not worth the effort. dplyr can also be fast, so it is a good choice if you want to speed things up, but don't quite need the scalability of data.table.
Many of the other solutions below do not require any additional packages. Some of them are even fairly fast on medium-large datasets. Their primary disadvantage is either one of metaphor or of flexibility. By metaphor I mean that it is a tool designed for something else being coerced to solve this particular type of problem in a 'clever' way. By flexibility I mean they lack the ability to solve as wide a range of similar problems or to easily produce tidy output.
Examples
base functions
tapply:
tapply(df$speed, df$dive, mean)
# dive1 dive2
# 0.5419921 0.5103974
aggregate:
aggregate takes in data.frames, outputs data.frames, and uses a formula interface.
aggregate( speed ~ dive, df, mean )
# dive speed
# 1 dive1 0.5790946
# 2 dive2 0.4864489
by:
In its most user-friendly form, it takes in vectors and applies a function to them. However, its output is not in a very manipulable form.:
res.by <- by(df$speed, df$dive, mean)
res.by
# df$dive: dive1
# [1] 0.5790946
# ---------------------------------------
# df$dive: dive2
# [1] 0.4864489
To get around this, for simple uses of by the as.data.frame method in the taRifx library works:
library(taRifx)
as.data.frame(res.by)
# IDX1 value
# 1 dive1 0.6736807
# 2 dive2 0.4051447
split:
As the name suggests, it performs only the "split" part of the split-apply-combine strategy. To make the rest work, I'll write a small function that uses sapply for apply-combine. sapply automatically simplifies the result as much as possible. In our case, that means a vector rather than a data.frame, since we've got only 1 dimension of results.
splitmean <- function(df) {
s <- split( df, df$dive)
sapply( s, function(x) mean(x$speed) )
}
splitmean(df)
# dive1 dive2
# 0.5790946 0.4864489
External packages
data.table:
library(data.table)
setDT(df)[ , .(mean_speed = mean(speed)), by = dive]
# dive mean_speed
# 1: dive1 0.5419921
# 2: dive2 0.5103974
dplyr:
library(dplyr)
group_by(df, dive) %>% summarize(m = mean(speed))
plyr (the pre-cursor of dplyr)
Here's what the official page has to say about plyr:
It’s already possible to do this with base R functions (like split and
the apply family of functions), but plyr makes it all a bit easier
with:
- totally consistent names, arguments and outputs
- convenient parallelisation through the
foreach package
- input from and output to data.frames, matrices and lists
- progress bars to keep track of long running operations
- built-in error recovery, and informative error messages
- labels that are maintained across all transformations
In other words, if you learn one tool for split-apply-combine manipulation it should be plyr.
library(plyr)
res.plyr <- ddply( df, .(dive), function(x) mean(x$speed) )
res.plyr
# dive V1
# 1 dive1 0.5790946
# 2 dive2 0.4864489
reshape2:
The reshape2 library is not designed with split-apply-combine as its primary focus. Instead, it uses a two-part melt/cast strategy to perform a wide variety of data reshaping tasks. However, since it allows an aggregation function it can be used for this problem. It would not be my first choice for split-apply-combine operations, but its reshaping capabilities are powerful and thus you should learn this package as well.
library(reshape2)
dcast( melt(df), variable ~ dive, mean)
# Using dive as id variables
# variable dive1 dive2
# 1 speed 0.5790946 0.4864489
Benchmarks
10 rows, 2 groups
library(microbenchmark)
m1 <- microbenchmark(
by( df$speed, df$dive, mean),
aggregate( speed ~ dive, df, mean ),
splitmean(df),
ddply( df, .(dive), function(x) mean(x$speed) ),
dcast( melt(df), variable ~ dive, mean),
dt[, mean(speed), by = dive],
summarize( group_by(df, dive), m = mean(speed) ),
summarize( group_by(dt, dive), m = mean(speed) )
)
> print(m1, signif = 3)
Unit: microseconds
expr min lq mean median uq max neval cld
by(df$speed, df$dive, mean) 302 325 343.9 342 362 396 100 b
aggregate(speed ~ dive, df, mean) 904 966 1012.1 1020 1060 1130 100 e
splitmean(df) 191 206 249.9 220 232 1670 100 a
ddply(df, .(dive), function(x) mean(x$speed)) 1220 1310 1358.1 1340 1380 2740 100 f
dcast(melt(df), variable ~ dive, mean) 2150 2330 2440.7 2430 2490 4010 100 h
dt[, mean(speed), by = dive] 599 629 667.1 659 704 771 100 c
summarize(group_by(df, dive), m = mean(speed)) 663 710 774.6 744 782 2140 100 d
summarize(group_by(dt, dive), m = mean(speed)) 1860 1960 2051.0 2020 2090 3430 100 g
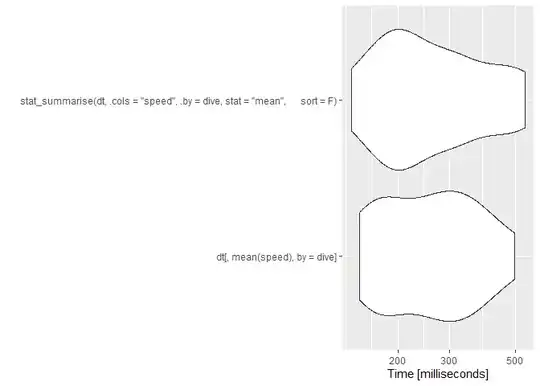
autoplot(m1)
As usual, data.table has a little more overhead so comes in about average for small datasets. These are microseconds, though, so the differences are trivial. Any of the approaches works fine here, and you should choose based on:
- What you're already familiar with or want to be familiar with (
plyr is always worth learning for its flexibility; data.table is worth learning if you plan to analyze huge datasets; by and aggregate and split are all base R functions and thus universally available)
- What output it returns (numeric, data.frame, or data.table -- the latter of which inherits from data.frame)
10 million rows, 10 groups
But what if we have a big dataset? Let's try 10^7 rows split over ten groups.
df <- data.frame(dive=factor(sample(letters[1:10],10^7,replace=TRUE)),speed=runif(10^7))
dt <- data.table(df)
setkey(dt,dive)
m2 <- microbenchmark(
by( df$speed, df$dive, mean),
aggregate( speed ~ dive, df, mean ),
splitmean(df),
ddply( df, .(dive), function(x) mean(x$speed) ),
dcast( melt(df), variable ~ dive, mean),
dt[,mean(speed),by=dive],
times=2
)
> print(m2, signif = 3)
Unit: milliseconds
expr min lq mean median uq max neval cld
by(df$speed, df$dive, mean) 720 770 799.1 791 816 958 100 d
aggregate(speed ~ dive, df, mean) 10900 11000 11027.0 11000 11100 11300 100 h
splitmean(df) 974 1040 1074.1 1060 1100 1280 100 e
ddply(df, .(dive), function(x) mean(x$speed)) 1050 1080 1110.4 1100 1130 1260 100 f
dcast(melt(df), variable ~ dive, mean) 2360 2450 2492.8 2490 2520 2620 100 g
dt[, mean(speed), by = dive] 119 120 126.2 120 122 212 100 a
summarize(group_by(df, dive), m = mean(speed)) 517 521 531.0 522 532 620 100 c
summarize(group_by(dt, dive), m = mean(speed)) 154 155 174.0 156 189 321 100 b
autoplot(m2)
Then data.table or dplyr using operating on data.tables is clearly the way to go. Certain approaches (aggregate and dcast) are beginning to look very slow.
10 million rows, 1,000 groups
If you have more groups, the difference becomes more pronounced. With 1,000 groups and the same 10^7 rows:
df <- data.frame(dive=factor(sample(seq(1000),10^7,replace=TRUE)),speed=runif(10^7))
dt <- data.table(df)
setkey(dt,dive)
# then run the same microbenchmark as above
print(m3, signif = 3)
Unit: milliseconds
expr min lq mean median uq max neval cld
by(df$speed, df$dive, mean) 776 791 816.2 810 828 925 100 b
aggregate(speed ~ dive, df, mean) 11200 11400 11460.2 11400 11500 12000 100 f
splitmean(df) 5940 6450 7562.4 7470 8370 11200 100 e
ddply(df, .(dive), function(x) mean(x$speed)) 1220 1250 1279.1 1280 1300 1440 100 c
dcast(melt(df), variable ~ dive, mean) 2110 2190 2267.8 2250 2290 2750 100 d
dt[, mean(speed), by = dive] 110 111 113.5 111 113 143 100 a
summarize(group_by(df, dive), m = mean(speed)) 625 630 637.1 633 644 701 100 b
summarize(group_by(dt, dive), m = mean(speed)) 129 130 137.3 131 142 213 100 a
autoplot(m3)
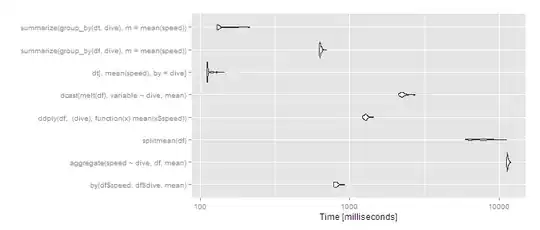
So data.table continues scaling well, and dplyr operating on a data.table also works well, with dplyr on data.frame close to an order of magnitude slower. The split/sapply strategy seems to scale poorly in the number of groups (meaning the split() is likely slow and the sapply is fast). by continues to be relatively efficient--at 5 seconds, it's definitely noticeable to the user but for a dataset this large still not unreasonable. Still, if you're routinely working with datasets of this size, data.table is clearly the way to go - 100% data.table for the best performance or dplyr with dplyr using data.table as a viable alternative.